
July 11, 2016
By: Ron Farber
The well-worn adage that “a picture is worth a thousand words” rings true when communicating the importance, content, and yes, the beauty that is uncovered as researchers explore how the brain works. Given that humans are wired to understand images faster and better than other forms of communication, brain research highlights the importance of scientific visualization
One of the demonstrations from ISC 2016 provided the opportunity for attendees to interact with the “Brayns” visualization application from the Swiss Blue Brain Project at the École Polytechnique Fédérale de Lausanne (EPFL). The demo highlighted that x86 multicore-based visualization can now be a viable, performant, and preferred path compared to GPU-based visualization.
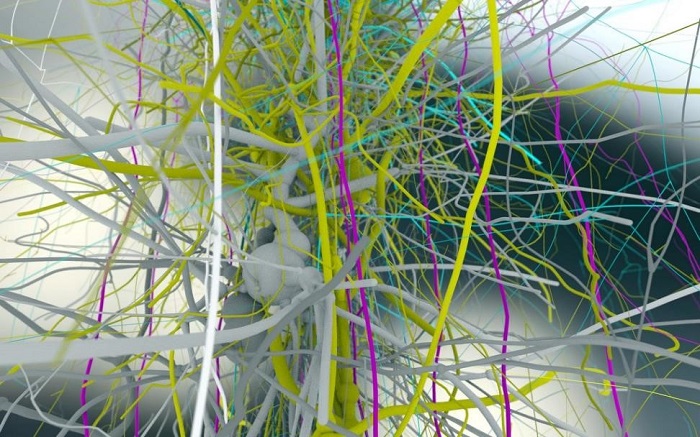

Figure 1: Even first in-silico models show the complexity and beauty of the brain
The Blue Brain demo was one of five live, interactive visualization demonstrations running on an Intel Scalable Systems Framework cluster using the new Intel 7210 Xeon Phi (“Knights Landing”) processors. Each processor provided 64-cores and were booted in self-hosted mode. The Brayns demonstration also took advantage of two open-source projects – OpenSWR, and OSPRay – that are being released as fully supported products in the software defined visualization (SDVis) initiative. The company’s visualization engineering group has already released the widely adopted open source Embree ray tracing library, as part of SDVis.
The multicore-based OSPRay ray-tracing engine was chosen by the Blue Brain Project as it is open-source, gives access to lots and lots of RAM, and runs everywhere. For example, OSPRay can be used on a MacBook Air for development, on workstations for the scientists, and in the data center using large memory fat-nodes. No GPU dependencies need be met on any of these platforms.
Succinctly, processor-based rendering lets the visualization team focus on what they are paid to do – generate images. Jim Jeffers, Engineering Manager & PE, Visualization Engineering at Intel, makes the point that, “SDVis matches the right quality and resolution visualization to the available hardware without imposing GPU dependencies.” The Blue Brain Project “Brayns” software is available for download from github: https://github.com/BlueBrain/Brayns.
The brain is arguably the most complex object we know, its biochemical and biophysical processes and structures span many spatial and temporal orders of magnitude. In addition to experimentation and theory, the study of the brain—one of the grand challenges of the 21st century—using modeling and simulation on computers has picked up momentum. Scientific visualization is an important part of those computer-based workflows. Using photo-realistic rendering is particularly suited to help scientists grasp the innate structures and discover salient patterns of activity.
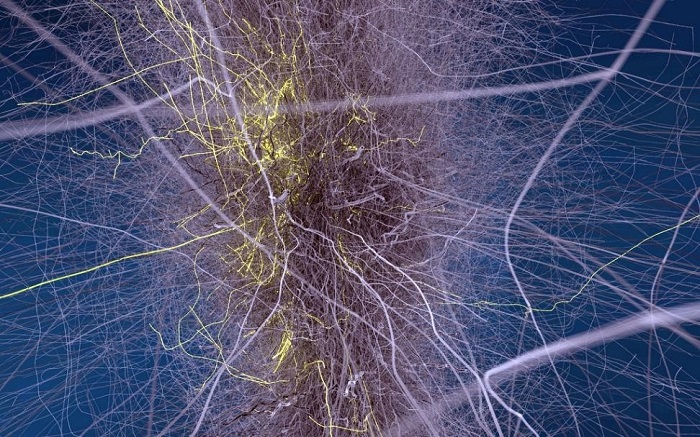

Figure 2: Visualizations provide detailed information to neuro scientists
The Blue Brain Project is using detailed modeling of brain tissue as a means to integrate multi-modal datasets. For example, the project has previously published in the scientific journal Cell a detailed reconstruction and simulation of the neocortical microcircuitry of a young rat. The work is introducing ray-tracing to improve previously possible visualizations. Before, during or after simulation, 3D visualization is a critical step for data analysis to enable insight, and specifically, ray-tracing can help to highlight areas of the circuits where cells touch each other and where synapses are being created. Using OSPRay’s ray tracing capability, using light, shadow, and depth of field effects that can simulate photo-realistic images, makes it much easier to visualize and therefore understand how the neurons function.

Figure 3: Highly detailed geometries showing intricate neural connectivity can be viewed at very fine detail
The ISC 2016 demo showed how computer and biological scientists take neural morphologies and reconstruct a piece of neocortex brain tissue, representing the cell structure using parametric geometry. Parametric equations are used to express the points that make up a geometric object such as the spheres and cylinders used in the neural visualization. Ray-tracing provides a natural way to work with these parametric descriptions so the image can be efficiently rendered in a much smaller amount of memory.
However, ‘smaller’ is a relative term as current visualizations can occur on a machine that contains less than a terabyte of RAM. Traditional raster-based rendering would have greatly increased the memory consumption as the convoluted shape of each neuron would require a mesh containing approximately 100,000 triangles per neuron. Scientists currently visualize biological networks containing 1,000 to 10,000 neurons.
The size and interactive nature of the Blue Brain Project demo is only possible now due to the significant increase in computing power made available over the past few years. Even a few years ago, interactive ray-tracing visualizations of this size was unthinkable. However, the memory savings and extraordinary quality of the images motivated that initially risky decision to use ray-tracing; a decision that has paid off handsomely and shows the performance available from the multicore-only hardware.
EPFL’s choice of open-source of the OSPRay project significantly reduced risk as it provided a proven ray-tracing engine, and which is also constantly being improved to keep abreast with the latest technology, such as Intel Xeon Phi processor and AVX-512 vectorization. A stable and established programming interface means that the newest OSPRay release can be dropped in to gain any new performance benefits, which further augments the visualization team’s ability do their work rather than delving down into the bowels of a complex software development effort. At the same time, having full source code access means the software developers can examine the code when needed to understand how the OSPRay engine works so they can press the limits of the technology, given the size and complexity of their visualization data sets both now and in the future.
The speed of the rendering depends on the quality and resolution of the destination image. Even with 1,000 to 10,000 neurons, the OSPRay engine delivers interactive performance for all datasets on a regular Intel Xeon processor, which can render images at 20-25 frames per second (FPS) on a 2k screen using 200-300 million geometries.
The Blue Brain Project demo represents a first-light viewing of images using the latest Intel Xeon Phi processors. The focus of the Blue Brain visualization effort over the past few years has been on functionality and proving that the interactive ray-tracing approach works. Initial impressions are that both the model and rendering applications contain both compute and memory limitations. The current belief is that memory bandwidth is the initial bottleneck. The expectation is that the significantly faster MCDRAM on the Intel Xeon Phi nodes should prove to be very helpful from a performance perspective, as will the dual AVX-512 vector units on each processor core.
The demo represented a risk to the EPFL team as they didn’t get access to the ISC 2016 cluster with Intel Xeon Phi processors until the day before the exhibition. However, they were willing to take the chance due to the open-source nature of the OSPRay source code and the consistency of the Intel Xeon Phi computing environment. As mentioned, the team had already built and run the OSPRay code on a variety of platforms from laptops to datacenter machines.
Similarly, Jim Jeffers’ team had four other live, interactive visualization demos that were seen for the first time running on the new Intel Xeon Phi processors at ISC 2016. Jeffers explained this makes their confidence concrete, “By preserving compatibility with previous generation products, Intel made the build-and-demo complex software on the new Intel Xeon Phi processors at a major trade show a low-risk proposition”.
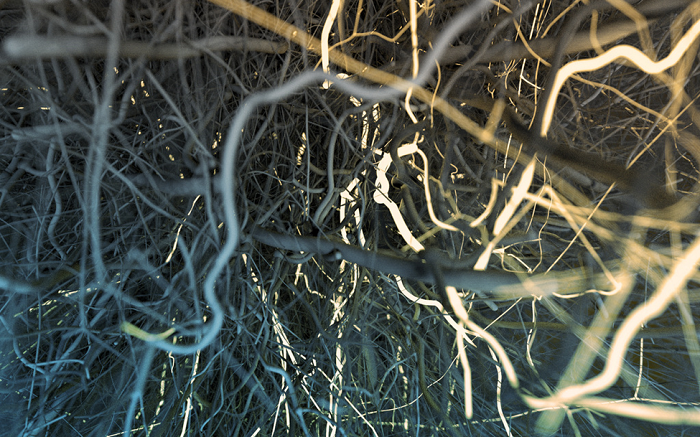
Figure 4: An image that uses Depth of Field (DOF)
Looking to the future, the Blue Brain visualization team hopes to start utilizing the multi-node capability of the OSPRay ray-tracing engine to increase the frame rate while also supporting higher resolution displays and display walls. Virtual reality is also a possibility. Utilizing lots of cores as well as multiple nodes means that the Blue Brain visualization team can also increase the number of rays to create even more beautiful images. Since scientists of the Blue Brain Project contribute to the large-scale effort of the European Human Brain Project, in the future this technology may help to convey the wonder and beauty that resides inside the human brain.
Figure 5: A plain image
Meanwhile, Jeffers is enthusiastic the future of the Intel supported SDVis effort as it will “provide new algorithms and datatypes and help realize the promise of in-situ visualization and full data distributed rendering.” Jeffers also noted that he is,“very excited about experimenting with 3D XPoint memory.”
-----
Rob Farber is a global technology consultant and author with an extensive background in scientific and commercial HPC plus a long history of working with national labs and corporations. He can be reached at info@techenablement.com.
